

Rice root–associated microbial community play an important role in plant nutrient acquisition, biomass production, and stress tolerance. Herein, root-associated community assembly was investigated under different phosphate input levels in phosphorus (P)-deficient paddy soil. Rice was grown in a long-term P-depleted paddy soil with 0 (P0), 50 (PL), or 200 (PH) mg P2O5 kg-1 application. DNA from root endophytes was isolated after 46 days, and PCR amplicons from archaea, bacteria, and fungi were sequenced by an Illumina Miseq PE300 platform, respectively. P application had no significant effect on rice root endophytic archaea, which were dominated by ammonia-oxidizing Candidatus Nitrososphaera. By contrast, rice root endophytic community structure of the bacteria and fungi was affected by soil P. Low P input increased endophytic bacterial diversity, whereas high P input increased rhizosphere fungi diversity. Bacillus and Pleosporales, associated with phosphate solubilization and P uptake, dominated in P0 and PH treatments, and Pseudomonas were more abundant in the PL treatment than in the P0 and PH treatments. Co-occurrence network analysis revealed a close interaction between endophytic bacteria and fungi. Soil P application affected both the rice root endosphere and soil rhizosphere microbial community and interaction between rice root endophytic bacteria, and fungi, especially species related to P cycling.
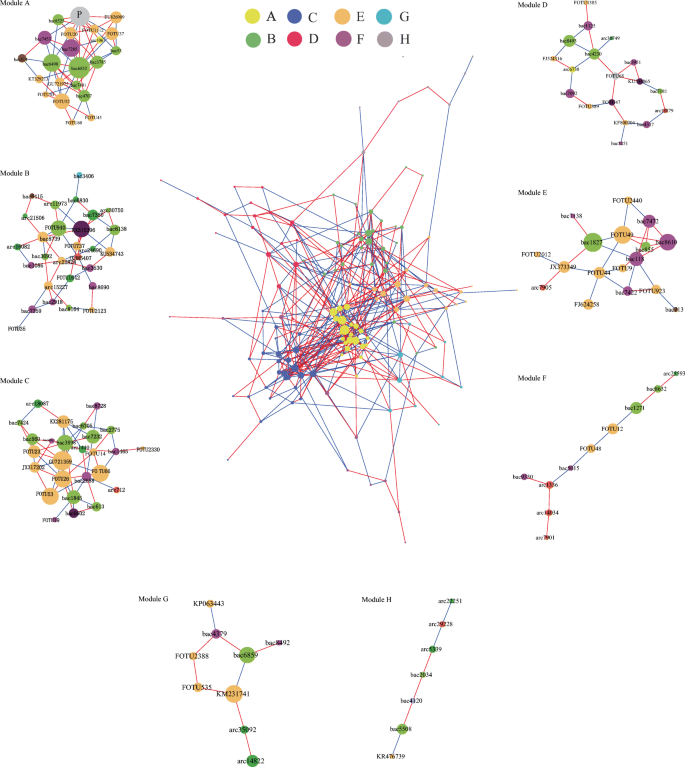
Co-occurrence network of rice root endophytes was generated by the greedy algorithm (GLay) clustering method. All nodes are grouped by modular pattern. The size of each node indicates its connections with other OTUs. Red and blue lines represent positive or negative correlations, respectively. Phylum information for modules A–H is presented in Fig S10